The health of cells in culture is critical to the success of your experiments. Have you ever been excited about the experimental results of a knock-down, drug treatment, or culture condition, only to realize later that the effects are skewed due to the amount of cell death that occurred in your samples? Measuring and comparing cell viability in your assays is important, whether it’s the data you’re pursuing or an important control in your experiment.
What is Cell Viability?
Cell viability refers to the proportion of healthy cells in a sample population. Measuring viability is essential to determine the physiological state of cells under experimental conditions. Cell viability is impacted by environmental factors, including alterations in cell culture parameters, exposure to pharmacological agents and growth factors, and in response to numerous disease states.
For example, cancerous cells exhibit enhanced viability, which enables them to proliferate abnormally and evade apoptosis, while neurodegenerative and other tissue degenerative disorders are characterized by progressive cell loss. Injury and infection also negatively impact cell survival, resulting in necrotic or apoptotic cell death. Therefore, the critical link between cellular health and disease highlights the necessity for assays that measure cell viability in different experimental contexts and model systems.
How to Measure Cell Viability
A variety of techniques have been developed to suit this purpose—that is, to measure cell viability across a range of experimental platforms. Viability assays can be grouped based on the underlying principle mechanism they investigate, including cell proliferation, plasma membrane integrity, metabolic activity, and mitochondrial function.
In addition to providing a snapshot of the overall health of a cellular population, assays can be tailored to also determine the method of cell death—through apoptosis or necrosis—when viability is compromised.
XTT Assay
The XTT assay is one example of a metabolic test to measure cell viability. In healthy cells, XTT is converted by mitochondrial enzymes into an orange formazan dye. Relative absorbance detected at 450 nanometers is then used to estimate the number of viable cells. One advantage of this method is its simplicity—as it does not require cell fixing, permeabilization, or secondary detection methods—and enables further examination of the cells following analysis.
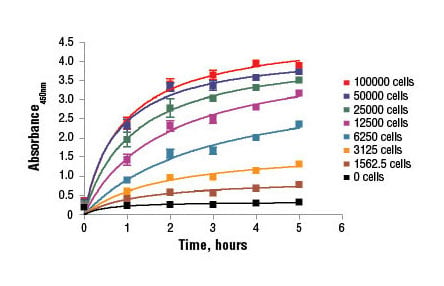 C2C12 cells were seeded at varying density in a 96-well plate and incubated overnight. The XTT Cell Viability Kit #9095 assay solution was added to the plate and cells were incubated. The absorbance at 450 nm was measured at 1.0, 2.0, 3.0, 4.0, and 5.0 hours.
C2C12 cells were seeded at varying density in a 96-well plate and incubated overnight. The XTT Cell Viability Kit #9095 assay solution was added to the plate and cells were incubated. The absorbance at 450 nm was measured at 1.0, 2.0, 3.0, 4.0, and 5.0 hours.
Resazurin Assay
Another example of a metabolic test for viability is the Resazurin assay. When added to cell cultures, the resazurin reagent is reduced to fluorescent resorufin by dehydrogenase enzymes in healthy cells, and the amount of resorufin is proportional to the viable cells in the sample. Similar to the XTT assay, cells used in the resazurin assay may be used for further downstream analysis.
Check out the Resazurin Cell Viability Kit Protocol for more details about how to use our Resazurin Cell Viability Kit #11884.
Live/Dead Cell Counting Assays
Live/dead cell counting assays are another popular method to assess cell viability. These tests use cell-permeable vital dyes, cell-impermeable dyes, or a combination of both to simultaneously label viable and dead cells within a population. For example, trypan blue is an azo dye that cannot penetrate cell membranes and is therefore excluded from healthy cells. However, dying cells with damaged membranes take up trypan blue, enabling the number of dead (blue) and living (colorless) cells to be counted using a hemocytometer.
TUNEL Assay
Methods like the terminal deoxynucleotidyl transferase dUTP nick end labeling (TUNEL) assay can also be used to detect cells undergoing programmed cell death. The assay uses a modified dUTP conjugated to a fluorophore to label the 3’OH end of fragmented DNA that is a hallmark of apoptotic cells. TUNEL assays are flexible as users can choose between different fluorophore options when multiplexing with antibody-based assays like immunohistochemistry, flow cytometry, and immunofluorescence.
.webp?width=341&height=337&name=25879_fig03_IHC-P_2624_20220203084000%20(1).webp)
Confocal analysis of paraffin-embedded human colon carcinoma using TUNEL Assay Kit #25879 (Fluorescence, 488 nm) (green) followed by staining with Cleaved Caspase-3 (Asp175) (5A1E) Rabbit mAb #9664 (red) and DAPI #4083 (blue).
CST offers three TUNEL kits that are robust, simple, and easy to use, and deliver results in cultured cells, tissue, and paraffin-embedded samples in one day.
Using IHC to Assay Cell Viability
Immunohistochemical (IHC) methods have also been developed to assay cell viability. This approach is commonly used to identify the temporal distribution of proteins and their expression patterns in healthy versus diseased tissue, as well as to identify areas of cells undergoing cell death while preserving the cytoarchitecture of intact tissue sections. In particular, markers of cellular proliferation, including Ki-67 or PCNA, or apoptosis, including cleaved caspase-3 and PARP, are commonly quantified by IHC.
Additional Resources:
- For a more in-depth discussion on the concepts and cell viability and applicable assays, please visit our webpage on cell death resources.
- To learn more about cell viability and the assays used to detect it, visit the following resource: Synopsis of Cell Proliferation, Metabolic Status, and Cell Death





/Binary_oasis_Blog%20Header.webp)
