Reagent validation and experiment reproducibility are critical for advancing research in any scientific discipline. Achieving reproducible data in (induced) pluripotent stem cell research presents particularly formidable challenges due to the biological complexity inherent to this research. The process of reprogramming cells and then subsequently directing them down different pathways of differentiation is fraught with technical challenges, not least of which is the remarkable complexity of events and interactions required to coax cells toward the desired state.
Essential Markers to Identify Induced Pluripotent Stem Cells (iPSC)
When programming cellular differentiation pathways, many points of confirmation are required throughout the process. How can researchers be sure that induced pluripotent stem cells (iPSC) actually become what was intended?
Researchers used to have to confirm pluripotency using functional assays, but can now use immunohistochemical markers to reliably assess pluripotency and to assess differentiation capacity (tri-lineage differentiation potential).
We caught up with Dr Harald Mikkers, Assistant Professor and co-head of the iPSC Core at Leiden University Medical Center, who works with iPSC every day and collaborated with us at CST to overcome some of these challenges.
The following is an account of the collaboration by Dr Mikkers:
"At the Leiden University Medical Center Human iPSC Hotel, we offer a cell reprogramming service to our clients (LUMC researchers as well as external commercial and non-profit parties). We isolate and expand cells derived from primary donor material such as skin, peripheral blood, and urine. The isolated primary cells are reprogrammed into induced pluripotent stem cells (iPSCs) with mRNA, episomal DNA, lentiviral vectors, or sendai virus.
The generated iPSC clones are evaluated for pluripotency status and functional pluripotency. To this end, we analyze the expression of pluripotent stem cell markers in the iPSC clones, and determine their capacity to differentiate into cells representing the three germ layers. The tri-lineage differentiation potential is qualitatively evaluated using fluorescent immunostaining.
To accelerate our workflow, we have developed a tri-lineage antibody kit consisting of directly conjugated CST antibodies against established markers of ectoderm (Figure 1), endoderm (Figure 2), and mesoderm (Figure 3). The conjugation (tier: 4 degrees of labeling) was performed by the custom conjugation service at CST. The success of conjugation was very high, and over 90% of the conjugations yielded antibodies that performed flawlessly in immunofluorescence microscopy experiments, at dilutions very similar to those recommended by CST for the respective unconjugated primary antibody.
Representative examples of the custom-conjugated antibodies are depicted below. In summary, conjugation of a defined set of primary antibodies permits cost- and time-effective characterization of pluripotency.
Conjugated Antibodies for Ectoderm, Endoderm & Mesoderm Characterization
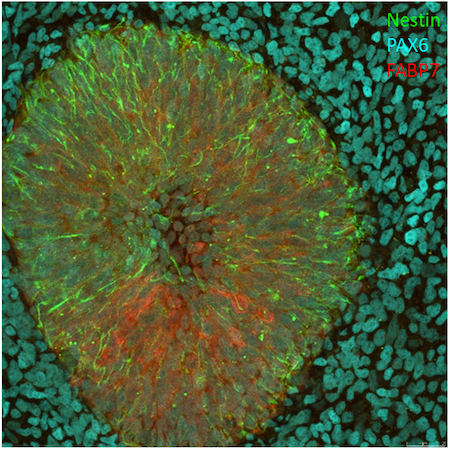 Figure 1. Ectoderm differentiation of LUMC0183iCTRL01. Antibodies used to detect ectoderm: Nestin (10C2) Mouse mAb (Alexa Fluor® 488 Conjugate) #35884, 42571 FABP7 (D8N3N) Rabbit mAb (Alexa Fluor® 555 Conjugate) and 84052 Pax6 (D3A9V) XP® Rabbit mAb (Alexa Fluor® 647 Conjugate).
Figure 1. Ectoderm differentiation of LUMC0183iCTRL01. Antibodies used to detect ectoderm: Nestin (10C2) Mouse mAb (Alexa Fluor® 488 Conjugate) #35884, 42571 FABP7 (D8N3N) Rabbit mAb (Alexa Fluor® 555 Conjugate) and 84052 Pax6 (D3A9V) XP® Rabbit mAb (Alexa Fluor® 647 Conjugate).
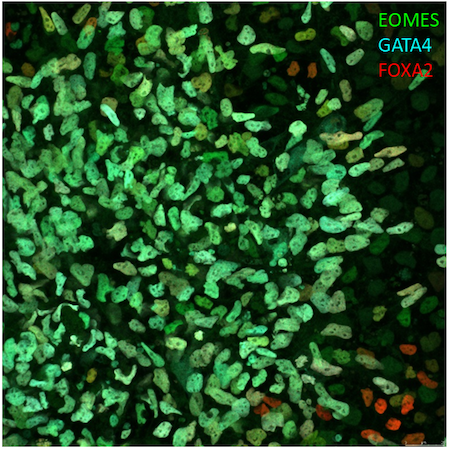 Figure 2. Endoderm differentiation of LUMC0183iCTRL01. Antibodies used to detect endoderm: EOMES (D8D1R) Rabbit mAb (Alexa Fluor® 488 Conjugate) #96823, FoxA2/HNF3β (D56D6) XP® Rabbit mAb (Alexa Fluor® 555 Conjugate) #50079 and GATA-4 (D3A3M) Rabbit mAb (Alexa Fluor® 647 Conjugate) #66309.
Figure 2. Endoderm differentiation of LUMC0183iCTRL01. Antibodies used to detect endoderm: EOMES (D8D1R) Rabbit mAb (Alexa Fluor® 488 Conjugate) #96823, FoxA2/HNF3β (D56D6) XP® Rabbit mAb (Alexa Fluor® 555 Conjugate) #50079 and GATA-4 (D3A3M) Rabbit mAb (Alexa Fluor® 647 Conjugate) #66309.
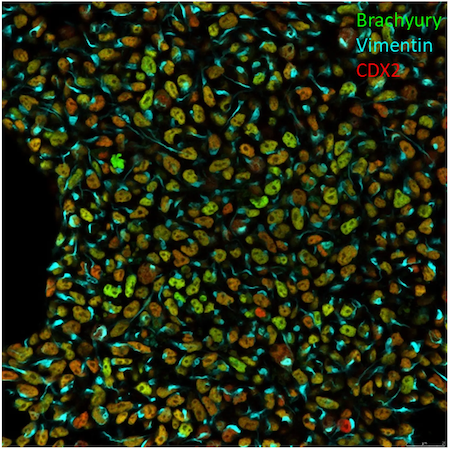 Figure 3. Mesoderm differentiation of LUMC0183iCTRL01. Antibodies used to detect mesoderm were: Brachyury (D2Z3J) Rabbit mAb (Alexa Fluor® 488 Conjugate) #9466, CDX2 (D11D10) Rabbit mAb (Alexa Fluor® 555 Conjugate) #84638 and Vimentin (D21H3) XP® Rabbit mAb (Alexa Fluor® 647 Conjugate) #9856
Figure 3. Mesoderm differentiation of LUMC0183iCTRL01. Antibodies used to detect mesoderm were: Brachyury (D2Z3J) Rabbit mAb (Alexa Fluor® 488 Conjugate) #9466, CDX2 (D11D10) Rabbit mAb (Alexa Fluor® 555 Conjugate) #84638 and Vimentin (D21H3) XP® Rabbit mAb (Alexa Fluor® 647 Conjugate) #9856
Learn More About CST Custom Conjugation Services
Perhaps the panel of antibodies we co-developed with Dr Mikkers could benefit your research, too. Maybe similar methods could be employed to validate cells in your experiments.
Could your research benefit from necessary, though difficult-to-produce reagents? Let us know about your research dilemma; we’re here to help!