In targeted protein degradation (TPD) drug discovery, confirming target ubiquitination is a pivotal step toward characterizing its degradation pathway. Yet for many scientists, getting those answers is anything but straightforward. While LC-MS is the most common method for assessing global protein ubiquitination, LC-MS cores are often booked solid, proteomics workflows can stretch weeks (or months), and data interpretation adds another layer of complexity. For teams under pressure to move projects forward, these bottlenecks can stall momentum at key decision points.
Whether you’re optimizing assay conditions at the bench or shepherding multiple projects through the pipeline, the challenge is the same: How to get quantitative, decision-ready ubiquitination data without relying on LC-MS‑based proteomics for every experiment?
First discovered in 2009, Tandem-Repeated Ubiquitin-Binding Entities (TUBEs) are a recombinant technology that can help overcome the sensitivity challenges of ubiquitination studies using antibody reagents alone, while helping teams avoid the LC-MS bottlenecks. By combining TUBEs with linkage‑, phospho‑, or target‑specific antibodies, you can build quantitative assays that answer questions such as “Is my target ubiquitinated?", “What linkage types are present?”, and “How does ubiquitination change with degrader dose over time?”
This post covers how to choose and deploy TUBEs‑based reagents alongside ubiquitin antibodies in western blot, ELISA+, TR-FRET, and other workflows, so you can match the assay to your specific TPD question, throughput needs, and sample constraints.
What Are Tandem-Repeated Ubiquitin-Binding Entities (TUBEs)?
Tandem Ubiquitin‑Binding Entities (TUBEs) are engineered proteins comprising a repeat of four ubiquitin-associated (UBA) domains arranged in series to provide exceptionally high apparent affinity for polyubiquitin chains. Functionally, they act as molecular traps that efficiently capture both K48- and K63-branched polyubiquitinated proteins, enabling sensitive detection of ubiquitination events that might be missed with standard ubiquitin antibodies alone.
 Figure 1. Tandem ubiquitin-binding entities (TUBEs) use four repeated ubiquitin-associated domains to bind polyubiquitin chains on target proteins for sensitive detection of ubiquitination.
Figure 1. Tandem ubiquitin-binding entities (TUBEs) use four repeated ubiquitin-associated domains to bind polyubiquitin chains on target proteins for sensitive detection of ubiquitination.
While traditional ubiquitin antibodies can struggle to efficiently immunoprecipitate (IP) diverse polyubiquitinated species, especially low‑abundance or heavily branched chains, TUBEs provide greater apparent affinity and more efficient capture due to their use of multiple UBA domains. Importantly, TUBEs selectively bind polyubiquitinated proteins rather than free ubiquitin, which improves specificity for ubiquitinated substrates in complex lysates. When combined with linkage-, phospho, and target-specific antibodies, this translates into stronger signal and better recovery of ubiquitinated proteins than branch‑specific or pan‑ubiquitin IP alone.
TUBEs can also help protect captured polyubiquitinated proteins from deubiquitinase activity during sample handling, preserving chains for downstream analysis.
Blog: Unlocking the Power of Ubiquitin: The Science Behind Targeted Protein Degradation
UBQLN1 vs RAD23A TUBES: What’s the Difference?
TUBEs reagents are derived from the ubiquitin receptor domains of UBQLN1 and RAD23A, two well-characterized ubiquitin-binding proteins. Each TUBE variant can bind both K48‑ and K63‑linked polyubiquitin chains, enabling broad capture of degradative and non‑degradative ubiquitination events. This pan‑branch binding profile makes TUBEs especially useful when the exact linkage pattern is unknown or may change with treatment.
To demonstrate the unique binding properties of UBQLN1- and RAD23A-derived TUBEs, CST researchers conducted a whole‑protein mass spectrometry experiment where polyubiquitinated proteins were captured using a Pan‑branch Ubiquitin TUBE‑RAD23A reagent compared with a Pan‑branch Ubiquitin TUBE‑UBQLN1 reagent.

Gary Kasof, Director of Product Design & Strategy |
“While UBQLN1- and RAD23A-based TUBEs each bind a vast array of polyubiquitinated proteins, target-specific preferences may exist. Using both can provide complementary performance depending on the substrate and linkage patterns in your system.” |
The total number of proteins detected after each capture showed that while most ubiquitinated proteins were common to both TUBE types, consistent with broad, pan‑branch enrichment, each reagent also pulled down a distinct subset of proteins that appeared enriched uniquely with either UBQLN1‑ or RAD23A‑based TUBEs.
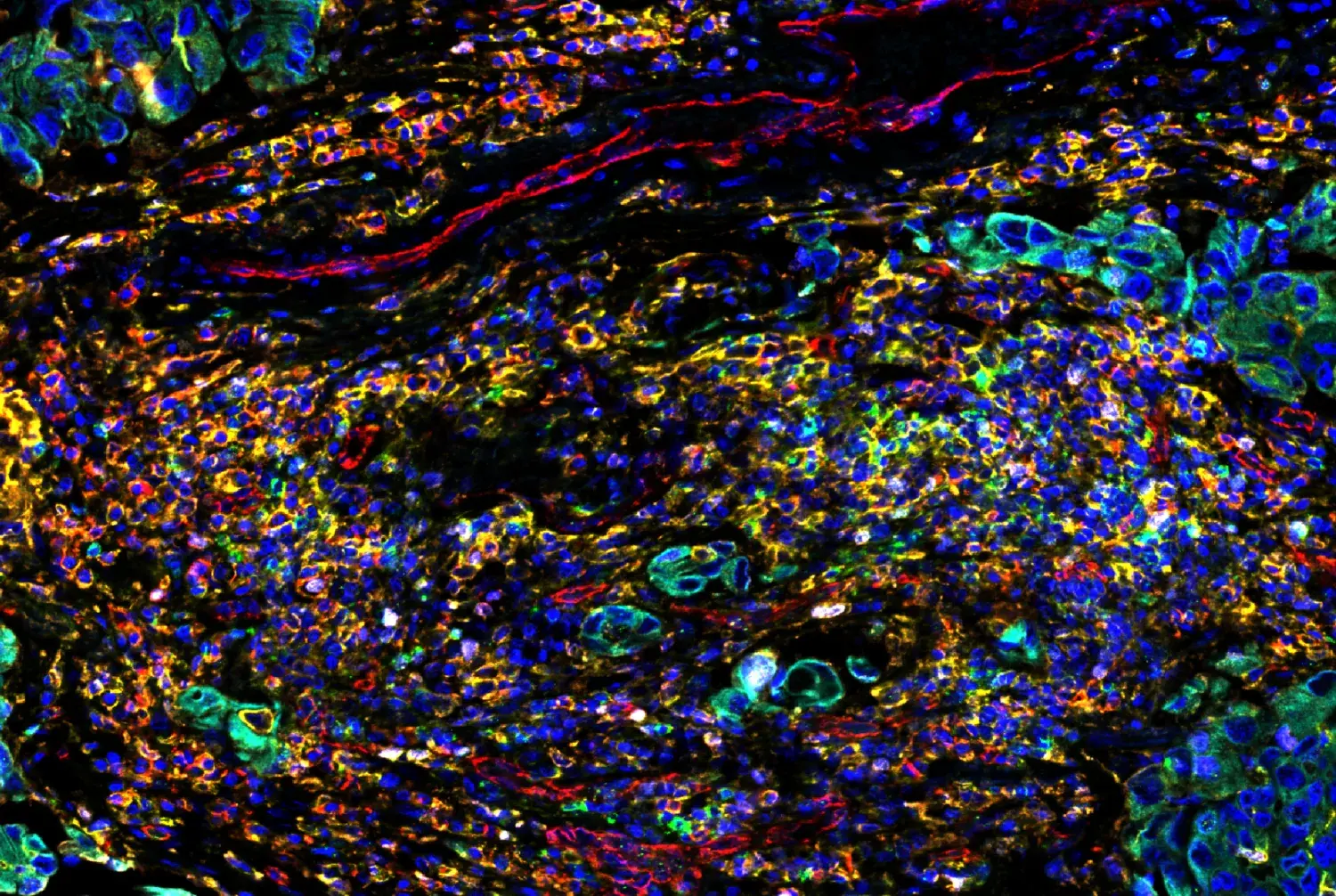
 Figure 2. Whole‑protein MS analysis of polyubiquitinated proteins captured with Pan‑branch Ubiquitin TUBE‑RAD23A (Magnetic Bead Conjugate) #64334 or Pan‑branch Ubiquitin TUBE‑UBQLN1 (Magnetic Bead Conjugate) #43658 from HCT 116 cells treated with or without multiple DUB inhibitors. Most proteins were captured by both TUBEs, indicating broad pan‑branch enrichment, but each TUBE also recovered a substantial set of apparently unique proteins, suggesting that using both reagents can provide more complete coverage of ubiquitinated substrates in complex samples.
Figure 2. Whole‑protein MS analysis of polyubiquitinated proteins captured with Pan‑branch Ubiquitin TUBE‑RAD23A (Magnetic Bead Conjugate) #64334 or Pan‑branch Ubiquitin TUBE‑UBQLN1 (Magnetic Bead Conjugate) #43658 from HCT 116 cells treated with or without multiple DUB inhibitors. Most proteins were captured by both TUBEs, indicating broad pan‑branch enrichment, but each TUBE also recovered a substantial set of apparently unique proteins, suggesting that using both reagents can provide more complete coverage of ubiquitinated substrates in complex samples.
These data suggest that while both tools provide deep coverage of polyubiquitinated species, using them in combination may offer more complete coverage of ubiquitination events, particularly in complex samples or when linkage patterns vary across targets and treatments.
Using TUBEs in Pair-Based Assays
Beyond simple pull-downs, TUBEs can be combined with a second detection reagent in a pair-based assay format, where one component captures ubiquitinated species and the other reports on a specific protein or post‑translational modification (PTM). In a typical setup, TUBEs first enrich polyubiquitinated proteins, and then a target‑specific antibody—for example, against your PROTAC target or a recruited E3 ligase—detects whether that protein is among the captured species, turning a generic ubiquitin capture into a target-focused readout. In degrader discovery, this modular capture‑and‑detect format allows teams to ask whether a given PROTAC or molecular glue engages its target and drives pathway‑relevant ubiquitination events.
Because capture and detection are modular, these pair‑based formats scale well to time courses, dose‑response studies, and panel screens across multiple cell models and degrader types.
|
TUBES-Based Assay Kits CST offers TUBEs immobilized on plates as capture reagents, creating “TUBE‑coated plate” configurations that support streamlined, wash‑based quantification of ubiquitinated species across many wells. |
|
Bead-Conjugated TUBES for Pull Downs
Bead-conjugated TUBEs (magnetic or Sepharose) enable efficient enrichment of polyubiquitinated proteins directly from cell or tissue lysates, followed by downstream detection of specific substrates via western blot. Compared with using ubiquitin or target antibodies alone, Pan‑branch TUBE‑UBQLN1 and TUBE‑RAD23A bead conjugates improve capture of low‑abundance, branched, or transiently ubiquitinated species that can otherwise fall below the detection threshold of antibody-only workflows.
This format is useful when you want to confirm ubiquitination of a specific target in western blot or use TUBEs as a capture reagent prior to proteomic LC‑MS analysis. In this way, bead‑conjugated TUBEs can complement other ubiquitination tools by enriching relevant polyubiquitinated substrates for either band‑level validation or deeper, site‑resolved proteomics.
Magnetic and Sepharose Bead-Conjugated TUBESReady-to-use enrichment reagents for pull‑down plus western blot workflows that can be combined with linkage‑, phospho‑, or target‑specific antibodies: |
|
For example, inflammatory stimuli such as TNF are known to increase ubiquitination of signaling proteins like RIPK1. Bead-conjugated TUBEs can efficiently pull down these polyubiquitinated forms (Figure 3), enabling downstream detection of RIPK1 and related pathway components using a blot. In this type of experiment, TUBEs capture polyubiquitinated substrates, and target‑specific antibodies such are then used in western blot to confirm which proteins are ubiquitinated under each condition.
 Figure 3. Left: Magnetic bead-conjugated TUBES used to isolate polyubiqinated proteins, which were then detected with Ubiquitin (E4I2J) Rabbit Monoclonal Antibody #43124. Does not bind to free ubiquitin. Right: Magnetic bead-conjugated TUBES used to isolate target-specific polyubiquitin from TNF-treated (+) and untreated cells using RIP (D94C12) Rabbit Monoclonal Antibody #3493.
Figure 3. Left: Magnetic bead-conjugated TUBES used to isolate polyubiqinated proteins, which were then detected with Ubiquitin (E4I2J) Rabbit Monoclonal Antibody #43124. Does not bind to free ubiquitin. Right: Magnetic bead-conjugated TUBES used to isolate target-specific polyubiquitin from TNF-treated (+) and untreated cells using RIP (D94C12) Rabbit Monoclonal Antibody #3493.
Because targeted substrates can be rapidly degraded after ubiquitination in PROTAC studies, short pretreatment with proteasome inhibitors is often used to stabilize ubiquitinated intermediates before TUBEs enrichment and detection, improving the chance of capturing transient species.
Biotin- or Europium-Conjugated TUBES for ELISA-Like and TR-FRET Assays
Biotinylated- or europium-conjugated TUBEs enable quantitative detection in higher-throughput, plate-based assay formats, making them well-suited for screening campaigns and detailed kinetic studies across many samples. In ELISA-like setups, biotinylated TUBEs capture polyubiquitinated species, while streptavidin-based detection and target- or PTM-specific antibodies provide readouts on the protein or modification of interest.
Biotin and Europium-Conjugated TUBESDesigned for ELISA+, TR‑FRET, and other homogeneous assay platforms where you need sensitive, plate-based detection of polyubiquitinated proteins: |
|
For example, CST’s biotinylated Pan-branch Ubiquitin TUBE-RAD23A #58553 can be used as a detection reagent in sandwich ELISA‑like formats to sensitively quantify ubiquitinated proteins across many samples and conditions. By pairing biotin‑compatible TUBEs with phospho‑specific antibodies against ubiquitin Ser65, researchers can monitor mitophagy‑associated signaling events, since PINK1‑dependent phosphorylation of ubiquitin at Ser65 is a well‑established marker and activator of Parkin‑mediated mitophagy.
In the example figures below, biotin‑ and europium‑conjugated TUBEs were used in ELISA‑like configurations to quantify pathway‑specific ubiquitination events, demonstrating how the same capture reagents can support both targeted and broader screening applications.
 |
 |
| Figure 4. Sandwich ELISA was performed using the Pan-branch Ubiquitin TUBE-UBQLN1 Assay Kit (Anti-rabbit IgG Secondary) #17452 with Phospho-Ubiquitin (Ser65) (E2J6T) Rabbit Monoclonal Antibody (1 µg/mL) as a detection antibody. This paired sandwich ELISA was used to detect stimulation of phospho-ubiquitin (Ser65) in valinomycin-treated 293 cells. The relationship between lysate protein concentration from untreated and valinomycin-treated 293 cells and the absorbance at 450 nm is shown in the left figure. The corresponding western blots using phospho-ubiquitin (Ser65) antibody (left panel) and ubiquitin antibody (right panel) are shown in the right figure. 293 cells were either left untreated or treated with valinomycin (1 µM) for 24 hr at 37°C, and then lysed. | Figure 5. Sandwich ELISA was performed using the Pan-branch Ubiquitin TUBE-UBQLN1 Assay Kit (Anti-rabbit IgG Secondary) #17452 with RIP (E8S7U) Rabbit Monoclonal Antibody (2 μg/mL) as a detection antibody. This paired sandwich ELISA was used to detect increased levels of ubiquitinated RIP protein in TNF-α-treated HeLa cells. The relationship between lysate protein concentration from untreated and TNF-α-treated HeLa cells and the absorbance at 450 nm is shown. HeLa cells were either left untreated or treated with TNF-α (20 ng/mL) for 5 min, and then lysed. |
When You Still Need LC‑MS
If your goal is unbiased, proteome‑wide mapping of ubiquitination sites, or you need to understand on‑ and off‑target ubiquitination networks for a degrader, LC‑MS is still the method of choice.
In a typical workflow, trypsin‑digested peptides are enriched using antibodies that recognize the K‑ε‑GG remnant left on lysines after ubiquitin (or ubiquitin‑like modifier) cleavage. This approach does not distinguish mono‑ from poly‑ubiquitination or resolve chain linkage types, but it provides a sensitive strategy for identifying global ubiquitination sites and quantifying changes across the proteome. Remnant‑motif antibodies (for example, PTMScan® HS Ubiquitin/SUMO Remnant Motif (K-epsilon-GG) Kit #59322) can also be used more narrowly to monitor ubiquitination of specific targets following PROTAC treatment and to complement total‑protein proteomics for deep mechanism‑of‑action studies.
Research Poster: Optimized Ubiquitin Remnant DIA Proteomics Enables High-Throughput Screening of PROTACs
In practice, many TPD teams use TUBEs‑based assays for hypothesis‑driven questions and screening, then reserve K‑ε‑GG LC‑MS for a smaller set of experiments where site‑level resolution and global network analysis are required.
Choosing the Right Ubiquination Detection Approach
Combining bead‑based and plate‑based TUBEs formats gives TPD teams a flexible toolkit: pull‑down assays for mechanistic, band‑level validation by western blot, and ELISA‑like or TR‑FRET configurations for higher‑throughput quantification of ubiquitination across many doses, time points, or cell lines. Selecting the right assay for your research depends on throughput, specificity, and the type of insight needed.
The table below maps common TPD questions to practical assay designs using TUBEs, antibodies, and LC‑MS.
| Research Question | Recommended Approach | When It Fits Best |
| Is my substrate ubiquitinated? | Capture of polyubiquitinated proteins with magnetic- or sepharose-conjugated TUBES, followed by western blot with a target-specific antibody. | Ideal for rapid, target‑focused confirmation of ubiquitination and for comparing conditions or doses without LC‑MS, especially when target-specific antibodies capable of immunoprecipitation are not available. |
| What type of ubiquitin linkage is present? | Immunoprecipitation with a target-specific antibody followed by western blot or IP using linkage‑specific ubiquitin antibodies (for example, K48- or K63-specific), combined with pan‑branch TUBEs for improved capture. | Useful when distinguishing K48‑, K63‑, or mixed‑linkage chains is critical for mechanism‑of‑action or pathway‑bias questions |
| How can I perform fast, sensitive, high‑throughput screening of ubiquitination across many conditions? | Biotinylated TUBEs in ELISA‑like or pair‑based assays, and europium‑based TR‑FRET setups for fast, sensitive high‑throughput detection. | Well‑suited for sensitive screening campaigns, SAR studies, and kinetic analyses across large panels of doses, time points, or cell models |
| What are the global, proteome‑wide changes? | Enrichment of ubiquitin remnants (for example, K‑ε‑GG peptides) followed by LC‑MS‑based proteomics | Remains a gold standard for unbiased, proteome‑wide mapping of ubiquitination sites and networks when time and instrumentation are available. |
Additional TPD Resources
- Resource Center: Tools for Proximity-Induced Targeted Protein Degradation
- Webinar: Targeted protein degradation: Controlling and assessing therapeutic targets to advance oncology drug discovery
- Speakers: Behnam Nabet, PhD (Fred Hutchinson Cancer Research Center), and Matthew Stokes, PhD (Cell Signaling Technology)
Select References
- Eladl O. Molecular glues and PROTACs in targeted protein degradation: mechanisms, advances, and therapeutic potential. Biochem Pharmacol. 2025;242(Pt 3):117297. doi:10.1016/j.bcp.2025.117297
- Hjerpe R, Aillet F, Lopitz-Otsoa F, Lang V, England P, Rodriguez MS. Efficient protection and isolation of ubiquitylated proteins using tandem ubiquitin-binding entities. EMBO Rep. 2009;10(11):1250-1258. doi:10.1038/embor.2009.192
- Snyder LB, Neklesa TK, Willard RR, et al. Preclinical Evaluation of Bavdegalutamide (ARV-110), a Novel PROteolysis TArgeting Chimera Androgen Receptor Degrader. Mol Cancer Ther. 2025;24(4):511-522. doi:10.1158/1535-7163.MCT-23-0655