Ubiquitin is a small but powerful protein that functions in nearly all cellular signaling pathways and is critical for regulating essential cellular processes. The attachment of one or more ubiquitin proteins, referred to as ubiquitination, can target proteins for degradation or modulate their activity, localization, or interactions—making ubiquitin a central regulator of protein quality control, signaling, and emerging therapeutic strategies such as targeted protein degradation (TPD).
Understanding how ubiquitin signaling is regulated is important for advancing TPD drug discovery, which has the potential to generate drugs with increased potency and enable the degradation of targets that were previously considered undruggable.
The Ubiquitination Pathway: How Proteins Get Tagged for Degradation
Ubiquitination is activated in a three-step, enzymatic process:
-
Ubiquitin Activation: First, ubiquitin is activated by an E1 enzyme, forming a thioester bond.
-
Ubiquitin Conjugation: The activated ubiquitin is transferred to a ubiquitin‑conjugating enzyme, E2.
-
Ubiquitin Ligation: Finally, the ubiquitin is transferred from E2 to specific substrates via E3 ubiquitin ligases.
The way ubiquitin molecules are connected to the target protein—the linkage or branch type—as well as post-translational modifications (PTMs) like phosphorylation, dramatically influence the biological function of ubiquitination and the fate of the protein substrate. Proteins can be either monoubiquitinated or polyubiquitinated, where polyubiquitin chains form either through specific lysine residues, with K48- and K63-branch linkages being the most abundant and widely studied, or with a head-to-tail configuration via the linear ubiquitin chain assembly complex (LUBAC).
- K48-branched polyubiquitination is recognized by the proteasome and typically leads to protein degradation.
- K63 branches and linear ubiquitination often regulate protein scaffolding and signaling pathways related to endocytosis, DNA damage control, and immune response.
Beyond K48 and K63, ubiquitin can form chains through other lysines (e.g., K11, K29, K33) and mixed chains, each of which signals a diverse set of cellular functions and outcomes.
Because different linkage patterns often correspond to distinct biological outcomes (for example, degradation vs signaling), linkage‑specific antibodies can help infer the functional consequences of ubiquitination on a given substrate. When paired with TUBEs (Tandem‑Repeated Ubiquitin‑Binding Entities) reagents, these linkage‑specific antibodies support both selective enrichment and linkage‑resolved detection of polyubiquitinated substrates in a given experiment.
Linkage-Specific Polyubiquitin Antibodies & Pan‑Branch TUBEsTo detect specific linkage types, branch‑specific ubiquitin antibodies can be used in western blot or other immunoassay formats: For detecting low-abundance samples or those with complex chains, TUBEs act as molecular traps that selectively enrich polyubiquitinated proteins while excluding free ubiquitin. They can be paired with linkage-specific antibodies and conjugated for use in bead‑based pull‑downs, plate‑based ELISA‑like formats, or homogeneous TR‑FRET assays to quantify ubiquitination without LC‑MS. |
|
Ubiquitin Regulation via Phosphorylation
Ubiquitin itself is also regulated by phosphorylation, which adds another layer of control. One of the most well-characterized events, ubiquitin phosphorylation at Ser65, is mediated by the kinase PINK1 following mitochondrial damage and plays a key role in the recognition of ubiquitinated proteins localized to the mitochondria for clearance by mitophagy. This process is critical for maintaining healthy mitochondria, and disruption of this pathway contributes to diseases such as Parkinson’s.

|
 |
| Western blot analysis of extracts from PC-3 cells, untreated or treated with carbonyl cyanide 3-chlorophenylhydrazone, using Phospho-Ubiquitin (Ser65) (E2J6T) Rabbit Monoclonal Antibody #62802 (upper), PINK1 (D8G3) Rabbit Monoclonal Antibody #6946 (middle), or GAPDH (D16H11) Rabbit Monoclonal Antibody #5174 (lower). |
TR-FRET assay was performed using Pan-branch Ubiquitin TUBE-UBQLN1 (trFluor™ Europium Cryptate Conjugate #75130 and custom conjugated Phospho-Ubiquitin (Ser65) (E2J6T) Rabbit Monoclonal Antibody #62802 (Alexa Fluor® 647 Conjugate) with titrations of valinomycin-treated and untreated 293 lysate. The relationship between the lysate protein concentration and the TR-FRET ratio is shown. The TR-FRET assay was run in a 384-well white plate and read on a BMG PHERAstar using laser excitation (337 nm). |
Because Ser65 phosphorylation on ubiquitin is tightly coupled to PINK1–Parkin pathway activation, it is widely used as a mitophagy marker. Antibodies such as CST’s Phospho‑Ubiquitin (Ser65) reagents can be used in western blot or ELISA+ assays to track PINK1‑Parkin–dependent mitophagy in cell models, providing an accessible readout of mitochondrial quality control. Reagents such as these enable precise monitoring of this important signaling event, without the need for more complex proteomics workflows.
Detecting Phosphorylated Ubiquitin at Ser65Phospho-specific antibodies and TUBEs targeting Ser65-phosphorylated ubiquitin enable precise monitoring of this important signaling event. These reagents allow researchers to follow how mitochondrial damage, PINK1 activity, and Parkin‑mediated mitophagy are connected in cell‑based models, without needing to immediately adopt more complex proteomics workflows. |
|
Targeted Protein Degradation: Turning Biology into Precision Therapy
The degradative role of ubiquitination has now been co-opted to create therapeutic opportunities for targeting specific proteins to be degraded (TPD). This emerging technology leverages chemically induced proximity (CIP) to join E3 ubiquitin ligases to disease-associated proteins of interest, leading to their degradation. There are two primary strategies:
-
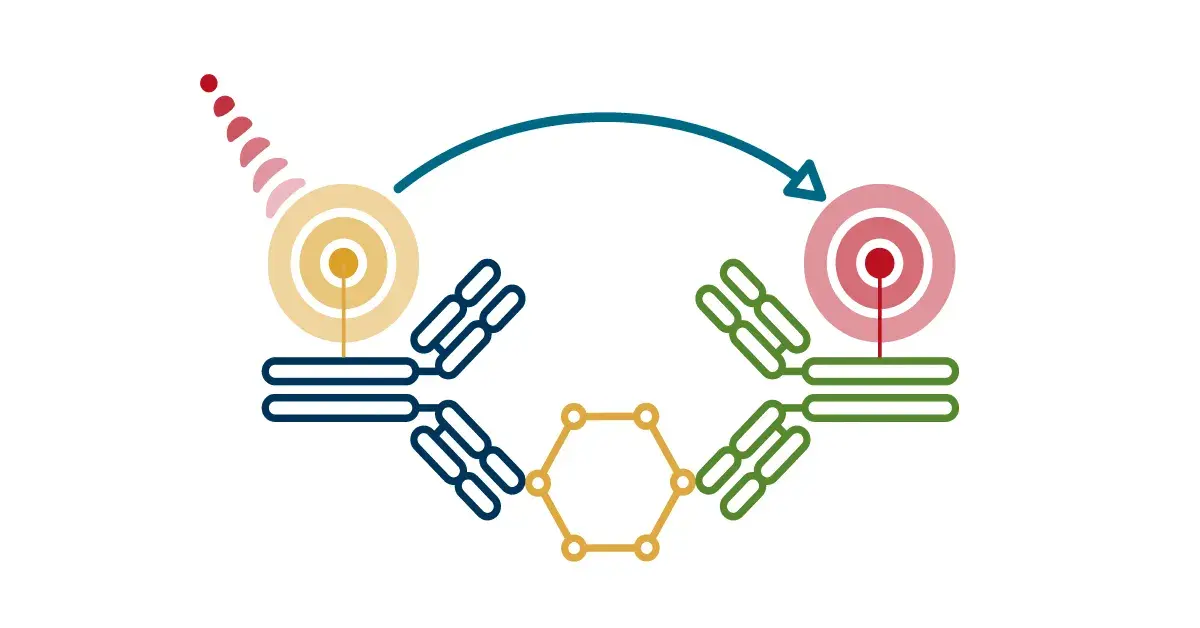
PROTACs (proteolysis-targeting chimeras): PROTACs are bifunctional molecules with two chemical binding domains: one that binds a protein of interest (POI) and another that binds an E3 ubiquitin ligase. By bringing the ligase and target into close proximity, PROTACs enable the cell’s own ubiquitin machinery to tag the POI for proteasomal degradation.
-
Molecular Glue Degraders: Molecular Glues utilize small molecules that stabilize the interaction between an E3 ubiquitin ligase and a POI, also resulting in ubiquitination and degradation.

Ubiquitination via a molecular glue degrader or PROTAC tags a target protein for recognition and degradation by the proteasome.
Although no drugs employing this approach have yet reached market approval as of early 2026, it is a rapidly growing area that holds promise to generate drugs with increased efficacy and potentially drug targets with weak binding pockets once thought to be undruggable. For example, previously undruggable targets such as mutated kinases (for example, KRAS), transcription factors (for example, STAT3), and protein scaffolds and complexes are now potentially druggable using TPD. Current clinical candidates, such as androgen receptor degraders for prostate cancer, illustrate how TPD can complement traditional small‑molecule inhibition by removing, rather than merely blocking, disease‑relevant proteins.
A range of platform-compatible, application-validated tools has emerged to answer critical questions about proximity-induced TPD throughout drug development. Depending on the question at hand, a combination of assays to profile ubiquitination can be used, including linkage‑ and phospho-specific antibodies, TUBEs‑based enrichment, or proteome‑wide K‑ε‑GG LC‑MS workflows, each of which offers a different balance of data, depth, and throughput.
Notably, because K‑ε‑GG–based enrichment detects the di‑glycine remnant left on lysines after trypsin digestion, it provides sensitive site‑level coverage of ubiquitination but does not distinguish mono‑ from poly‑ubiquitination or resolve chain linkage types. Branch-specific antibodies can help make this distinction, but standard ubiquitin antibodies may fail to efficiently pull down low‑stoichiometry or transiently ubiquitinated species. TUBEs, which are derived from ubiquitin receptors such as UBQLN1 and RAD23A, act as molecular traps that selectively enrich polyubiquitinated proteins while excluding free ubiquitin, improving both recovery and specificity for downstream analysis.
To explore how these tools are applied in TPD workflows—such as monitoring target ubiquitination, pathway activation, and on/off‑target effects—see the companion blog on assay selection and TUBEs‑based detection approaches:
Additional TPD Resources
-
Resource Center: Tools for Proximity-Induced Targeted Protein Degradation
- Webinar: Targeted protein degradation: Controlling and assessing therapeutic targets to advance oncology drug discovery
- Speakers: Behnam Nabet, PhD (Fred Hutchinson Cancer Research Center), and Matthew Stokes, PhD (Cell Signaling Technology)
Select References
- Eladl O. Molecular glues and PROTACs in targeted protein degradation: mechanisms, advances, and therapeutic potential. Biochem Pharmacol. 2025;242(Pt 3):117297. doi:10.1016/j.bcp.2025.117297
- Hjerpe R, Aillet F, Lopitz-Otsoa F, Lang V, England P, Rodriguez MS. Efficient protection and isolation of ubiquitylated proteins using tandem ubiquitin-binding entities. EMBO Rep. 2009;10(11):1250-1258. doi:10.1038/embor.2009.192