The average human brain contains over 100 billion cells, which is nearly the same number as there are stars in the Milky Way, making it, truly, an astronomical number! Together, these brain cells drive our ability to think, remember, feel emotions, and engage with the world. These cells truly make us who we are. The primary cell in the brain is the neuron, a highly polarized cell with molecularly specialized arbors, which initiates and drives information transfer and ultimately drives brain function. However, other non-neuronal cells, including oligodendrocytes, microglia, and astrocytes, play contributing if not critical roles in maintaining proper neuronal function. They also play crucial roles in development and disease.
Staining of these brain cell types is critical not only to identify brain cell structure but also to identify specific cells in the brain. Multiplexing techniques and probes enable neuroscientists to measure multiple molecular targets, simultaneously, in a single experiment. This technique facilitates molecular profiling of individual cells in the context of a network or in tissue, thereby providing a snapshot of cellular function.
Why multiplex the brain?
Cells in the brain include neurons, astrocytes, microglia, and oligodendrocytes. These cells have unique molecular signatures. You may be familiar with some of these cellular signatures, including NeuN (neurons), GFAP (astrocytes), Myelin Basic Protein, Olig2 (Oligodendrocytes), and Iba1 (macrophages/microglia). Probes to detect these molecular signatures can include complementary RNA probes that detect the transcript using techniques like in situ hybridization.
Although antibody-based methods can lack some of the sensitivity and specificity that one can approach with RNA-based methods, the detection of proteins using antibodies holds several advantages. For one, antibody-based detection can reveal the localization of a particular protein, suggesting a specific function. For example, the astrocyte protein GFAP is a cytoskeletal protein that reveals astrocytic morphologies.
 Immunofluorescent (IF) analysis of mouse hippocampus using GFAP (E4L7M) XP® Rabbit mAb #80788 (green), showing the astrocytic morphology.
Immunofluorescent (IF) analysis of mouse hippocampus using GFAP (E4L7M) XP® Rabbit mAb #80788 (green), showing the astrocytic morphology.
In neurons, NeuN is localized to the nucleus, as would be expected for a transcription factor. Staining with antibodies like MAP2 and Neurofilament light chain (NfL) reveal highly arborized neurons by labeling dendrites and axons, respectively. Deeper interrogation of subcellular proteins, especially in neurons, can reveal additional insights into the structure and function of cells.
Many different types of proteins are involved in the formation, maintenance, and plasticity of synapses, the central unit by which neurons communicate. Proteins may include cell adhesion molecules, adaptor proteins, and scaffolding proteins. Synaptic proteins such as synaptophysin and PSD95 (pre- and post-synaptic proteins, respectively) are enriched at neuronal synapses. Using multiplexing techniques, researchers can examine the protein’s co-localization of their favorite protein with a known synaptic protein. For example, colocalization of their protein of interest with a postsynaptic protein like PSD95 can suggest a postsynaptic function.
Multiplexing Methods
Researchers use multiple methods to effectively multiplex their staining experiments. One of the simplest ways to multiplex is to use primary antibodies derived from various host species and isotypes (rabbit IgG, mouse IgG1, rat IgG2a, etc.) and use fluorophore-coupled secondary antibodies specific to the host primary antibody used.
Another method is to directly conjugate a primary antibody with a specific fluorophore or enzyme. Often, antibodies are not well suited for this approach—e.g., available antibodies are from the same host species, or they are not conjugated with the appropriate detection modality. To get around this, researchers can use a TSA stripping protocol that involves sequential rounds of labeling and stripping with fluorophore-conjugated tyramide.
Blog: CST® Antibodies Validated on the Leica Microsystems Cell DIVE Multiplexed Imaging Solution
All of these approaches have their advantages and disadvantages. However, the brain can be uniquely sensitive and complex, and detection of low-abundance targets can be difficult. More sophisticated methods are emerging. For example, DNA oligo-labeled antibodies with subsequent signal amplification enables simultaneous protein staining far beyond 3-4 proteins stained compared to more traditional methods.
Using Multiplexing to Study Alzheimer’s Disease
The development and continued evolution of multiplexing technologies coincides with important research questions emerging in neuroscience, particularly in areas like Alzheimer’s disease. One of the pathological hallmarks of Alzheimer’s disease is amyloid plaques, which correlate with clinical-behavioral pathology as well as neuronal pathology. Recently, the contribution of non-neuronal cells in the pathology of this neurodegenerative disease has come under intense investigation. Much of this work is driven by new technologies like single cell RNAseq, which allows transcriptomic analysis at a single cell level. As a result, the field has only recently begun to appreciate the molecular diversity of microglia.
 |
Explore the interactive Molecular & Cellular Biology of Alzheimer’s Disease Pathway Diagram to see key protein targets and associated CST products. |
Microglia are the resident macrophages of the brain that are mobilized to detect and clear debris, including damaged neurons and plaques. Analysis of these cells in normal and diseased conditions revealed differential transcriptomic profiles, suggesting that distinct microglia play a key role in disease progression.1,2
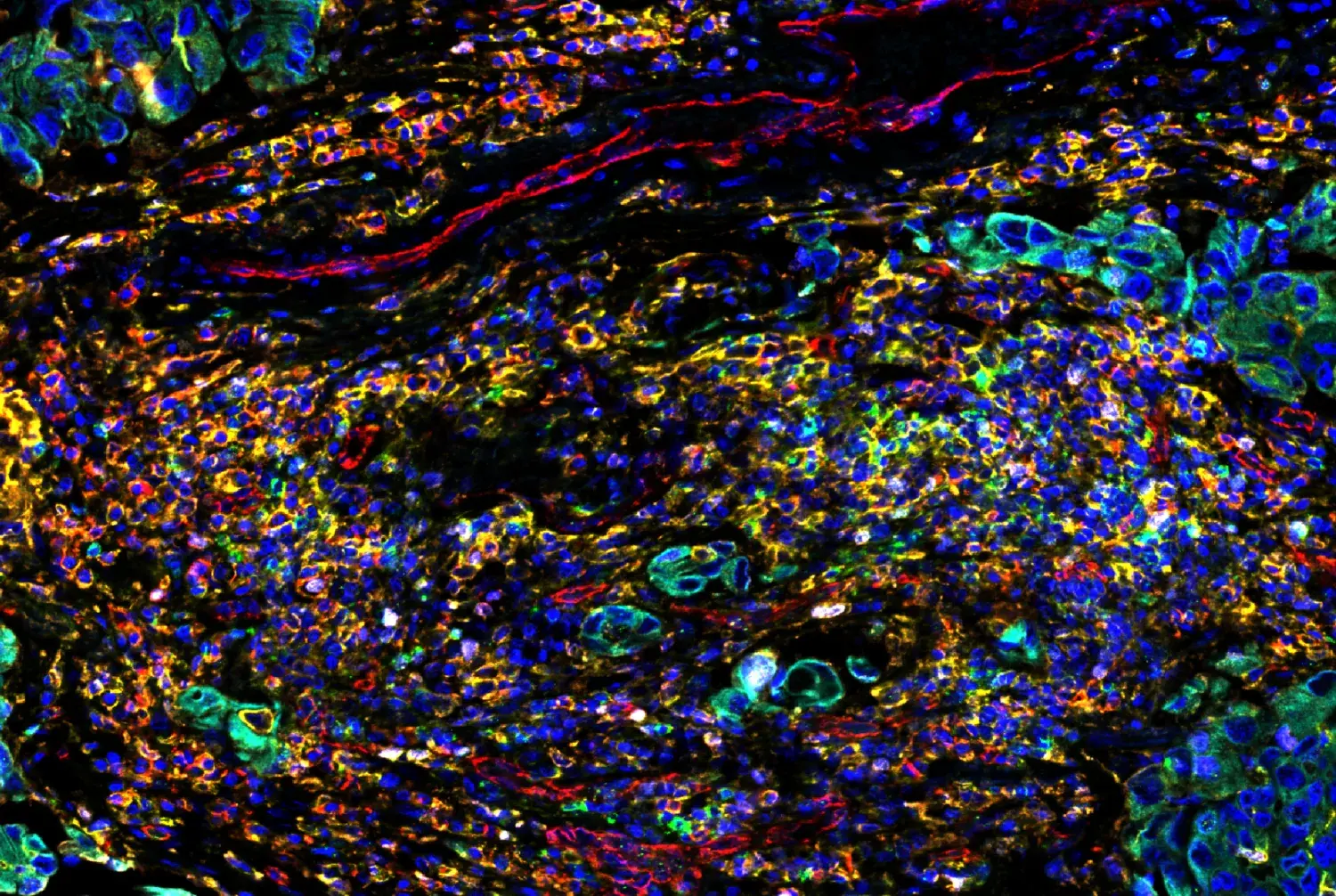
Although transcriptomic analysis has revealed the microglia complexity that exists in the brain, proteins are the functional currency of cells. A current challenge in the field is to use multiplexing techniques to investigate specific microglia in the context of disease. For example, researchers reported distinct transcriptomic signatures that correlate with disease, so-called disease-associated microglia (DAM’s).1 GPNMB, a gene that encodes a glycoprotein that functions as a negative regulator of inflammation, is highly upregulated in AD mouse models. Using antibodies to stain GPNMB, researchers can probe microglia-expressing GPNMB in the context of other brain cells and disease pathology. In Figure 1, microglia-expressing GPNMB protein (yellow) surrounds amyloid plaques (blue) suggesting that the protein is playing a direct role in disease progression. As the transcriptomics data suggested, these microglia are distinct DAMs as only a subset of IBA1-positive microglia staging for GPNMB. Identifying DAM by their proteomic profile within the context of tissue and disease enables researchers to understand the unique role microglia might be playing in disease progression.
Confocal IF analysis of brain from an amyloid mouse model of Alzheimer’s disease. Sections were labeled with Iba1/AIF-1 (E4O4W) XP Rabbit mAb (Alexa Fluor® 647 Conjugate) (Red), HS1 (D5A9) XP Rabbit mAb (Rodent Specific) (Alexa Fluor 488 Conjugate) (Green), GPNMB (E7U1Z) Rabbit mAb (Yellow), and methoxy XO4 (Blue). Images kindly provided by Dr Simone Briochi of Dr Marco Colonna's lab (Washington University) and used with permission.
As new technologies evolve, staining techniques using multiplexed probes that are antibody- or transcript-based will be increasingly important to understand the molecular and cellular mechanisms of complex processes as well as disease. In the brain, several challenges remain. For example, substantial auto-fluorescence is observed in the aged human brain, complicating fluorescent-based examination. Amplification methods, such oligo-based PCR, may improve signal-to-noise ratio that may make analysis of this tissue easier. Currently, most researchers use labor-intensive processes to slice the tissue into thin sections to facilitate probe penetration and imaging. However, whole tissue clearing techniques and advancing imaging techniques like light-sheet fluorescence microscopy will greatly facilitate multiscale staining and imaging. Such advancements will greatly facilitate the ability to make sense of the myriad of questions that remain in the brain sciences.
Looking for more multiplexing resources? The Neuronal and Glial Cell Marker Atlas is a useful guide for identifying cell type markers:
 |
Explore the interactive Neuronal and Glial Cell Marker Atlas to see key protein targets and associated CST products. |
Additional Resources
- Blog: Spatial Protoemics: Multiplexed Methods for Studying the Spatial Biology of the Brain
- Blog: Pros and Cons of Different Multiplexing Techniques for IHC and IF
- Blog: Strategies for Successful Multiplex IF Experiments Using Conjugated Antibodies
Select References:
- Keren-Shaul H, Spinrad A, Weiner A, et al. A Unique Microglia Type Associated with Restricting Development of Alzheimer's Disease. Cell. 2017;169(7):1276-1290.e17. doi:10.1016/j.cell.2017.05.018
- Mathys H, Davila-Velderrain J, Peng Z, et al. Single-cell transcriptomic analysis of Alzheimer's disease [published correction appears in Nature. 2019 Jun 17;:]. Nature. 2019;570(7761):332-337. doi:10.1038/s41586-019-1195-2
21-EMG-51658