Understanding the microenvironment of disease is critical. Targeted cancer therapies may not be effective depending on the combination of cell types in the tumor. Likewise, an immune cell invasion in pancreatic tissue could indicate the progression of type 1 diabetes. For years, this complex landscape could only be witnessed in single-cell snapshots by flow cytometry or limited tissue staining. Imaging Mass Cytometry (IMC) now offers a panorama of this dynamic world.
The Imaging Mass Cytometry Process
IMC enables imaging of up to 40 protein markers on both the cellular and tissue levels. Tissue samples are labeled with antibodies, each conjugated to a different rare-earth metal isotope. The sample is then placed within a chamber where a laser degrades the antibody, releasing the isotope into an adjacent time-of-flight mass cytometer (CyTOF) for ionization and identification by an associated mass-to-charge ratio. Compilation of these data by marker type and location results in a multidimensional tissue profile (Figure 1).
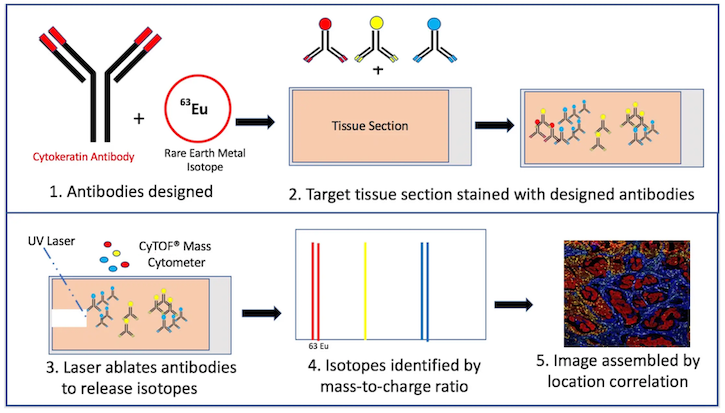
Figure 1. Mass cytometry process by Kris Hargraves. Assembled image courtesy of Dr Noel de Miranda.
Dr Charlotte Giesen at the University of Zürich, Switzerland first reported this method for tissue analysis in 2014. IMC enabled imaging of 32 proteins and their associated post-translational modifications at a cellular resolution of 1 µm in breast cancer tissue samples.1
IMC imaging of these samples often correlated with the pathologist’s classification of breast cancer subtype by traditional tissue staining. Exceptions, however, demonstrated greater tumor heterogeneity. At least one case classified as triple negative for the receptors ER, PR, and HER2 demonstrated subpopulations of HER2+ cells. This result indicates that targeted therapies against the HER2 receptor might be a viable treatment option for that patient.
Benefits of IMC
Colocalized detection of several markers greatly extends the boundaries of traditional tissue staining by immunofluorescence. Rare-earth metal isotopes are not naturally found in cells, eliminating intrinsic background concerns. Marker detection by mass also enables greater target specificity, unlike immunofluorescent markers that often overlap in their emission profile.
In a recent collaboration, Dr Noel de Miranda and Dr Frits Koning, both of the Leiden University Medical Center, used IMC to detect foreign antigen exposure within the human fetal intestine.2
“It is a great technique to investigate interactions between different cell subsets in the same tissue,” Dr de Miranda told Cell Signaling Technology.
Challenges of IMC
The technology is not without challenges. Antibodies labeled with rare-metal isotopes are currently not commercially available for many targets. This fact, plus the need for a robust signal in the appropriate tissue sample preparation, often requires researchers to generate their own antibody panels.
“It is a laborious and onerous task to build a panel of approximately 40 antibodies,” Dr. de Miranda said. “You want to use antibodies that have been thoroughly validated so that the majority of them work right off the bat.”
Carrier-Free Mass Cytometry Antibodies from CST
Dr de Miranda said that tools from CST, such as validated carrier-free antibodies, greatly assist his IMC work (Figure 2) and emphasized that target specificity of the primary antibody in the sample is critical for success.
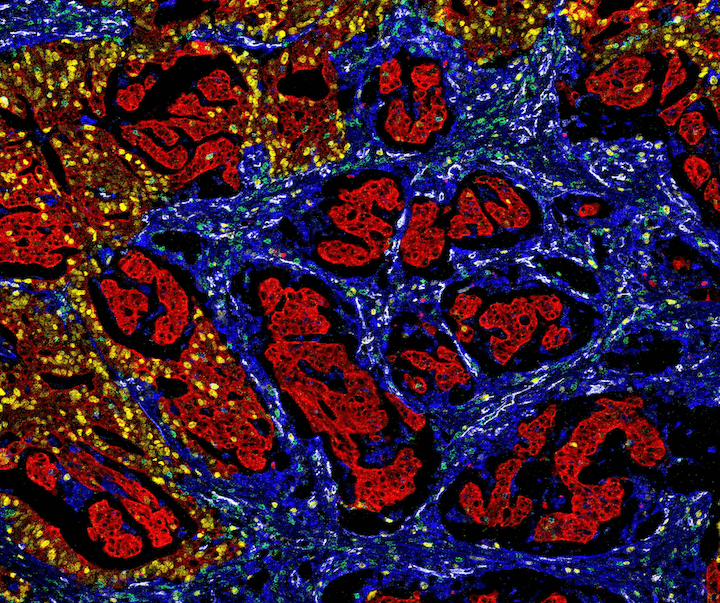 Figure 2. Mass cytometry image of colorectal cancer tissue with the following protein targets: Pan-Keratin (C11) Mouse mAb #4545 and a mouse Anti-pan cytokeratin antibody (red), Vimentin (D21H3) XP® Rabbit mAb #5741 (blue), Ki-67 (8D5) Mouse mAb #9449 (yellow), CD31 (PECAM-1) (89C2) Mouse mAb #3528 (white), and CD3ε (D7A6E™) XP® Rabbit mAb #85061 (green). Image courtesy of Dr Noel de Miranda.
Figure 2. Mass cytometry image of colorectal cancer tissue with the following protein targets: Pan-Keratin (C11) Mouse mAb #4545 and a mouse Anti-pan cytokeratin antibody (red), Vimentin (D21H3) XP® Rabbit mAb #5741 (blue), Ki-67 (8D5) Mouse mAb #9449 (yellow), CD31 (PECAM-1) (89C2) Mouse mAb #3528 (white), and CD3ε (D7A6E™) XP® Rabbit mAb #85061 (green). Image courtesy of Dr Noel de Miranda.
Dr de Miranda stated that support from CST "has been fundamental to advance our research, and we keep working together on the development of new technological approaches.” IMC continues to unlock the formerly hidden world of the tissue microenvironment.
Select References
- Giesen C, Wang HA, Schapiro D, et al. Highly multiplexed imaging of tumor tissues with subcellular resolution by mass cytometry. Nat Methods. 2014;11(4):417-422. doi:10.1038/nmeth.2869
- Li N, van Unen V, Abdelaal T, et al. Memory CD4+ T cells are generated in the human fetal intestine. Nat Immunol. 2019;20(3):301-312. doi:10.1038/s41590-018-0294-9



/Binary_oasis_Blog%20Header.webp)