Enabled by recent advances in sequencing, mass spectrometry, and fluorescence microscopy, spatial biology—the interrogation of genes and proteins in their original location within a tissue—is helping unravel the mysteries of cellular neighborhoods and their influence on the behavior of individual cells. This drastic increase in the volume and variety of data that can be acquired from a single tissue sample is revealing insights about how complex and constantly evolving relationships between cellular proximity, organization, and the timing of signaling events can affect disease progression and therapeutic response.
How can researchers more effectively leverage spatial biology to rapidly answer complex biological questions? SignalStar Multiplex IHC, a new spatial biology technology from CST, drastically decreases the time from assay design to data acquisition using flexible, ready-to-go antibody panels for multiplex immunohistochemistry (mIHC) experiments. With validated protocols and optimized reagents for interrogating the expression of up to eight targets in formalin-fixed, paraffin-embedded (FFPE) tissues, researchers can generate critical spatial biology data 70% faster from sample to imaging than other mIHC methods.
Why choose multiplex immunohistochemistry for spatial biology research?
Multiplex IHC is a powerful and versatile spatial biology tool that allows researchers to answer a range of questions by labeling multiple proteins in a single tissue sample. The technology is ideal for maximizing the data acquired from limited FFPE tissue, and can provide a detailed map of the spatial distribution and expression modification patterns of multiple targets (Figure 1).
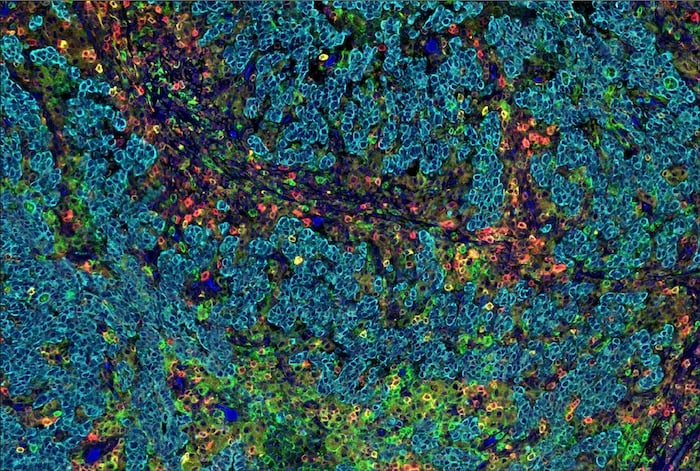
Figure 1. SignalStar Multiplex IHC assay analysis of multiple proteins in human gastric adenocarcinoma. Staining was performed on the BOND RX Fully Automated Research Stainer by Leica Biosystems.
While RNA analysis can provide insights about gene expression, it’s unable to capture post-translational modifications (PTMs) such as phosphorylation, glycosylation, and acetylation. Multiplex IHC overcomes these limitations by simultaneously identifying multiple PTMs within their spatial context, helping uncover biomarker coexpression patterns and providing a more comprehensive understanding of the underlying biology.
Spatial biology applications leveraging mIHC have proven particularly useful for understanding the tumor microenvironment (TME) and, more specifically, how tissue organization and the identity of cells in close proximity to the tumor affect disease progression. Spatial biology allows cancer researchers to ask questions such as:
- Does a tumor demonstrate heterogeneity in gene and protein expression compared to surrounding tissue?
- Which immune cells are present in the TME? Do the same immune cells exist in the surrounding tissue, or are they unique to the tumor?
- Are immune cells in an exhausted state, or are they primed to respond to the cancer?
These questions, which can be answered using mIHC in combination with various spatial biology techniques, are relevant for predicting a patient’s response to cancer immunotherapies—and, therefore, can help enable the future of personalized medicine.
However, many current spatial biology imaging techniques limit forward progress due to the time-consuming methods needed for assay design and antibody panel development, optimization, validation, and analysis. Some traditional mIHC assays require the use of primary antibodies developed in different species, each of which then necessitates its own unique secondary antibody detection system. This can limit the number of markers that can be labeled at once, and can lead to issues of cross-reactivity and background noise, making it difficult to obtain reliable and accurate results.
Additionally, challenges with alternative staining methods, such as tyramide signal amplification (TSA), include:
- Long protocol times: Each antibody needs to be added individually, and all reagents must be titrated with respect to each other.
- Antibody cycling: The need for antibody cycling can result in epitope loss and tissue degradation, reducing data quality.
- Order optimization: The order in which antibodies are added and imaged needs to be optimized, a lengthy process that requires re-optimization each time the antibody panel changes.
- Tyrosine deposits: With each antibody and tyramide staining cycle, the availability of tyrosine residues may be reduced, which can inhibit additional fluorophore deposition from subsequent antibodies, resulting in loss of signal.
To overcome these limitations, many researchers spend countless hours designing, optimizing, and validating antibody panels, a process that can take weeks to months. When a new target is identified or research goals shift, this optimization and validation process begins again. Additional challenges occur when researchers leverage antibodies that aren’t validated for the applications in which they will be used, or attempt to perform signal amplifications.
An assay is only as good as the antibody used, and ineffective reagents can waste precious tissue samples.
The SignalStar Multiplex IHC technology eliminates these hurdles by providing customizable, flexible, and highly validated antibody panels that work right out of the box.
A Spatial Biology Revolution: How does the SignalStar technology work?
SignalStar Multiplex IHC is a new technology for spatial biology research that uses high-throughput, mid-plex IHC assays to label up to eight targets in FFPE tissues simultaneously with flexible, highly validated antibody panels. The optimized, ready-to-go panels can be used to interrogate cellular presence, location, function, and biomarker coexpression patterns.
The technology uses a combination of oligonucleotides and fluorophores to amplify the antibody signal, allowing for the detection of targets, even with low expression levels. Using the SignalStar Multiplex IHC Panel Builder, researchers can select targets from a menu of IHC-validated antibodies and confidently analyze limited, precious FFPE samples. And, as research changes and evolves or new targets are identified, SignalStar antibodies and fluorophores are interchangeable, allowing researchers to switch out targets and quickly redesign panels without the need for additional panel or protocol optimization.
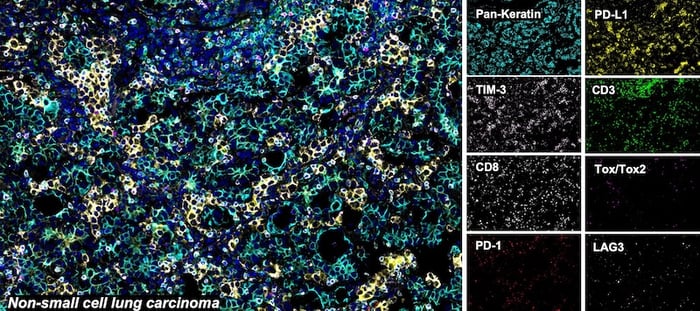
SignalStar® multiplex immunohistochemical analysis of paraffin-embedded human non-small cell lung carcinoma using CD3ε (D7A6E) & CO-0001-488 SignalStar® Oligo-Antibody Pair #92856 (green), PD-L1 (E1L3N) & CO-0005-594 SignalStar® Oligo-Antibody Pair #28249 (yellow), PD-1 (Intracellular Domain) (D4W2J) & CO-0008-647 SignalStar® Oligo-Antibody Pair #56837 (red), Pan-Keratin (C11) & CO-0003-750 SignalStar® Oligo-Antibody Pair #97227 (cyan), TIM-3 (D5D5R) & CO-00010-488 SignalStar® Oligo-Antibody Pair #81365 (light pink), LAG3 (D2G4O) & CO-0026-594 SignalStar® Oligo-Antibody Pair #34308 (orange), Tox/Tox2 (E6I3Q) & CO-0016-647 SignalStar® Oligo-Antibody Pair #67341 (magenta), CD8α (D8A8Y) & CO-0004-750 SignalStar® Oligo-Antibody Pair #62750 (white), and ProLong Gold Antifade Reagent with DAPI #8961 (blue). All fluorophores have been assigned a pseudocolor, as indicated. Staining was performed on the BOND RX autostainer by Leica Biosystems.
By leveraging the technology, spatial biology data can be achieved in days, not weeks or months, up to 70% faster than with other, traditional mIHC methods.
To learn more about SignalStar and how to speed your spatial biology research, download the brochure:
Alexandra Foley, Scientific Marketing Writer and CST Blog Manager, contributed to writing this post.23-ETC-78401






/Binary_oasis_Blog%20Header.webp)
